Open Science Portal
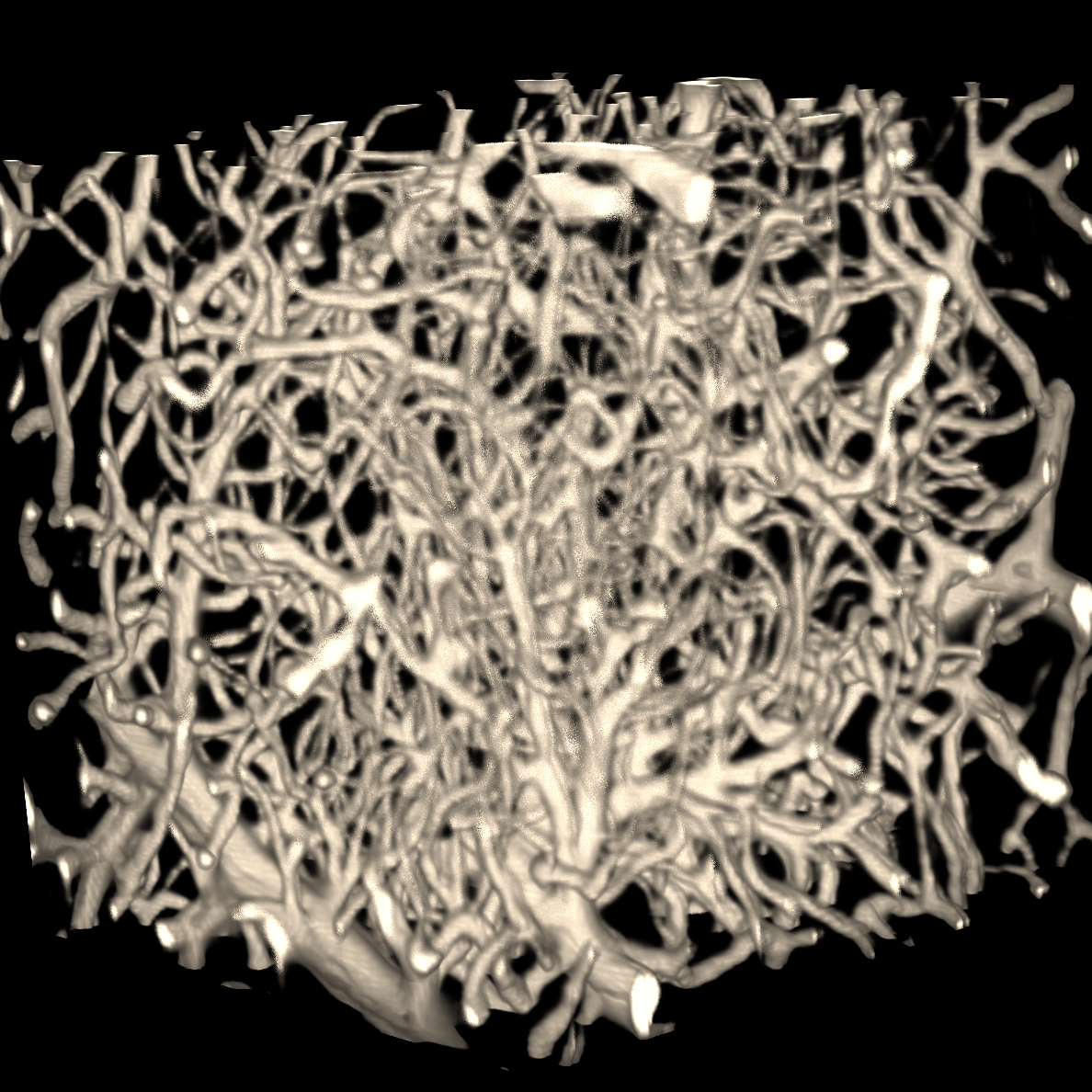
Our group is fully commited to open science, from developing/contributing to open source softwares to building public datasets for AI research as well as open access publications. We believe open science can lead to impactful big science. Currently, our group has a special focus on building AI-based biomedical image analysis methods/tools/datasets for light-sheet fluorescence microscopy (LSFM) imaging. Meanwhile, on this page, you can also find other open science works our group members built before, other than LSFM, as well as a curated list of open science tools developed by other researchers in the community that we would highly recommend.
Open source softwares
Open datasets
Other open science works from our group members
Other open source tools from the community (we would highly recommend)
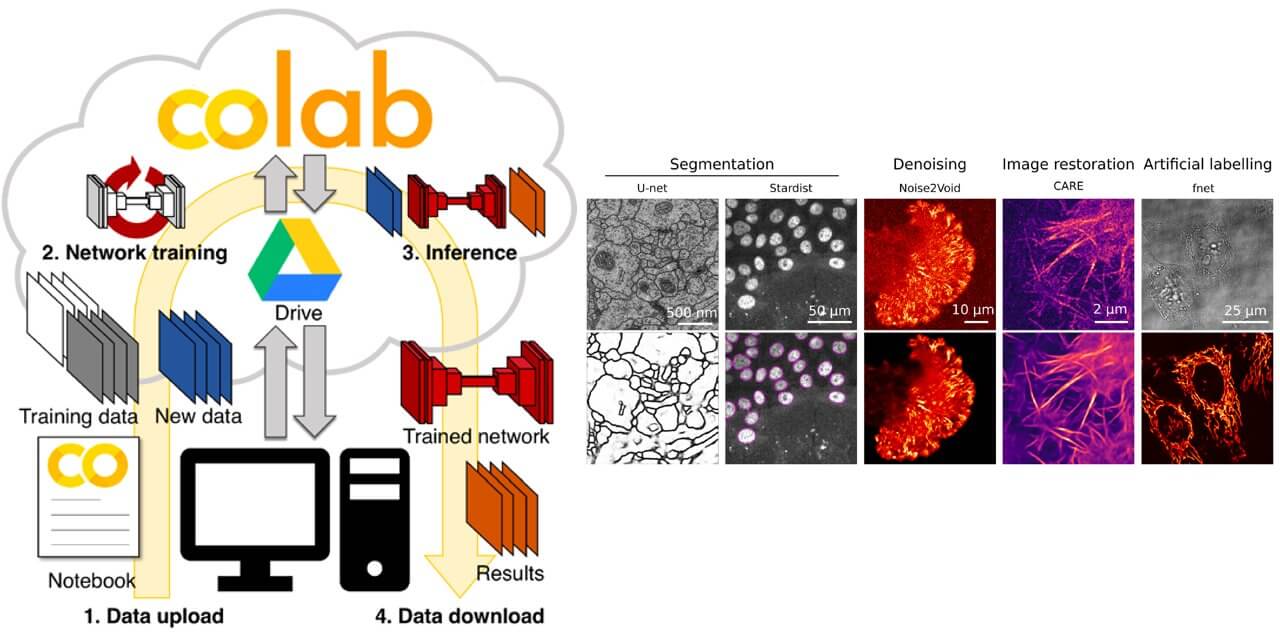
A collection of self-explanatory Jupyter Notebooks for Google Colab that features an easy-to-use graphical user interface. A good resource for learning deep learning applications in microscopy image analysis.